A Guide to Graph Representation Learning

In recent years, there has been a significant amount of research activity in the graph representation learning domain. These learning methods help in analyzing abstract graph structures in information networks and improve the performances of state-of-the-art machine learning solutions for real-world applications, such as social recommendations, targeted advertising, user search, etc. This article provides a comprehensive introduction to the graph representation learning domain, including common terminologies, deterministic and stochastic modeling techniques, taxonomy, evaluation methods, and applications.
Learning Embeddings from Graph Data
A graph is a fundamental data structure that has a set of vertices (also called nodes, points, or entities) and edges (also called relationships, or connections) that are used to model relationships among a collection of objects. Graph structures are ubiquitous throughout the natural and social sciences. For example, to model a social network as a graph, we can use nodes to represent individuals and edges as an indication that the two individuals are friends. Similarly in e-commerce businesses like Amazon, users and items can be nodes and the buying and selling relationships can be the edges.
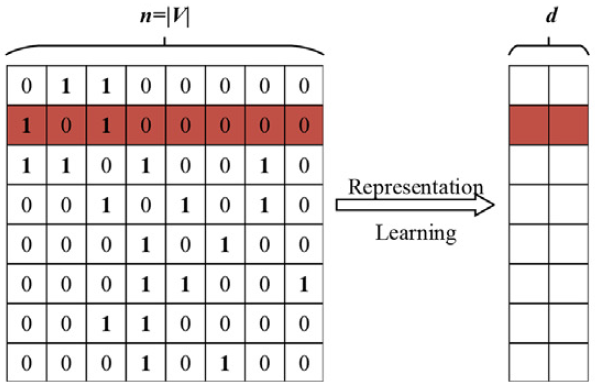
The process of extracting insights (also called network mining) from graph data has applications in a wide range of domains, like social computing, biology, recommender systems, language modeling, etc. Graph representation learning has been a very active research area for the last few years. To facilitate efficient graph data analysis, these representation learning algorithms encode the graph to a low-dimensional vector space. The goal of these learned vectors (also called embeddings) is to accurately capture the structure and features of complex high-dimensional graphs.
Graph embeddings are low-dimensional representations that minimize redundancy and noise in the original data. They also help in avoiding the need to devise complex algorithms that apply directly to the original graph. These vectors are later used as features in various downstream tasks, such as node classification, link prediction, clustering, recommendations, etc. Hence the quality of representation of these learned vectors is especially important. These embeddings must maintain network topology, vertex features, labels, and other auxiliary information in low-dimensional space such that the nodes that were similar in the original graph also have similar embeddings.
Note that the embeddings can be generated for nodes, edges, subgraphs, or even entire graphs. However, utilizing node embeddings for representing graphs is a more common and well-studied problem. After these vertex embeddings are learned, any vector-based machine learning algorithm can be efficiently applied to perform network analytic tasks in the new representation space.
Leveraging Network Structure
A lot of the approaches for graph representation explicitly leverage the connections between nodes. One popular idea is to exploit homophily, which is defined as the tendency for nodes to share attributes with their neighbors. For example, people tend to form friendships with other people who share the same interests or demographics. So the representation methods try to assign similar values to the neighboring nodes. Similarly, there are more concepts like structural equivalence and heterophily that can be exploited. We will look at some of the commonly used similarity measures in a later section.
Challenges
The goal of the representation learning process is to find a way to incorporate graph structural information into algorithms like machine learning. The central problem is that there is no straightforward way to encode all the high-dimensional and non-Euclidian graph information into feature vectors. The learning process also has to deal with the following challenges.
Preserve Structure
As stated earlier, we want similar vertices in the original graph to be represented similarly in the learned vector space. In the graph structure, the similarity between nodes is not only reflected in the local community structure but also in the global network structure. Therefore the learning process must preserve both local and global structures simultaneously.
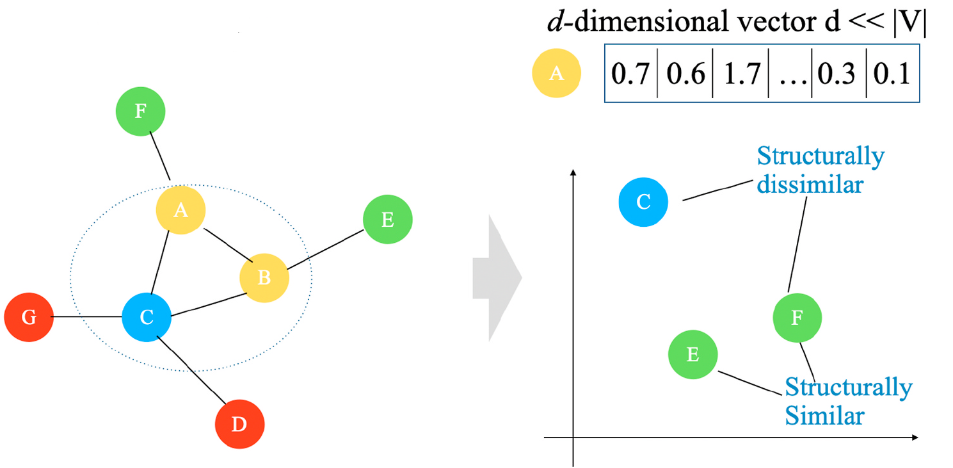
Preserve Content
Graph vertices often have rich attribute content that provides attribute-level node similarity information. The learning process becomes more complex when it utilizes network structure along with node attributes due to the heterogeneity of the two information sources.
Data Sparsity
In real-world networks, both the network structure and node attributes are missing due to privacy or legal restrictions. Nodes might not be explicitly connected due to this data sparsity, which increases the difficulty of learning structure-level node relatedness.
Scalability
Enterprise networks often have millions of billions of nodes. Therefore it is necessary to design efficient learning algorithms that avoid large computational and storage costs but also guarantee effectiveness.
Some Useful Definitions
Graph
Formally, a graph $G$ can be defined as a tuple $G = (V, E)$ where $V$ is the set of nodes/vertices and $E$ is the set of edges. A graph can be directed or undirected. In a directed graph (also called di-graph), each edge $e_k = (v_i, v_j)$ has a direction from starting vertex $v_i$ to the ending vertex $v_j$.
Homogeneous and Heterogeneous Graphs
Let’s assume that $T_V$ denotes the set of node types and $T_E$ denotes the set of edge types. A homogeneous graph is a graph with only one node type and one edge type (i.e. $|T_V| = 1$ and $|T_E| = 1$). In a heterogeneous graph $|T_V| + |T_E| > 2$.
Dynamic Graphs
Real-world networks are always evolving, such that new nodes or edges may be added or existing ones might be dropped between adjacent snapshots. Such a graph is defined as a series of snapshots $G = {G_1, G_2, … G_n }$ where $n$ is the number of snapshots.
Knowledge Graphs
In a multi-relational graph $G = (V, E)$, each edge is defined as a tuple $e_k = (v_i, \tau, v_j)$, where $\tau$ represents a particular relationship $\tau \in T$ between the two nodes. Although there is no single unifying definition of knowledge graphs, these multi-relational graphs can also be called knowledge graphs because the tuple represents a particular “fact” that holds between the two nodes.
Vertex Proximities
First-order proximity: Nodes that are connected with an edge have first-order proximity. Edge weights represent proximity measures such that the higher weight indicates more similarity between the two connected nodes1.
Second-order proximity: The second-order proximity represents the 2-step relations between each pair of vertices, and is defined as the number of common neighbors shared by the two vertices2.
Higher-order proximity: The higher-order proximity between two vertices $v$ and $u$ can be defined as the $k$-step ($k \ge 3$) transition probability from vertex $v$ to vertex $u$.
K-partite Graphs
In many real-world graphs, we have groups of nodes where edges are allowed between groups but not between nodes of the same group. For example, a bi-partite graph has two groups of nodes where nodes of one group can only connect to nodes of the other group. In k-partite graphs, “partite” refers to the partitions of node groups, and “k” refers to the number of those partitions. The following figure shows a tri-partite graph.
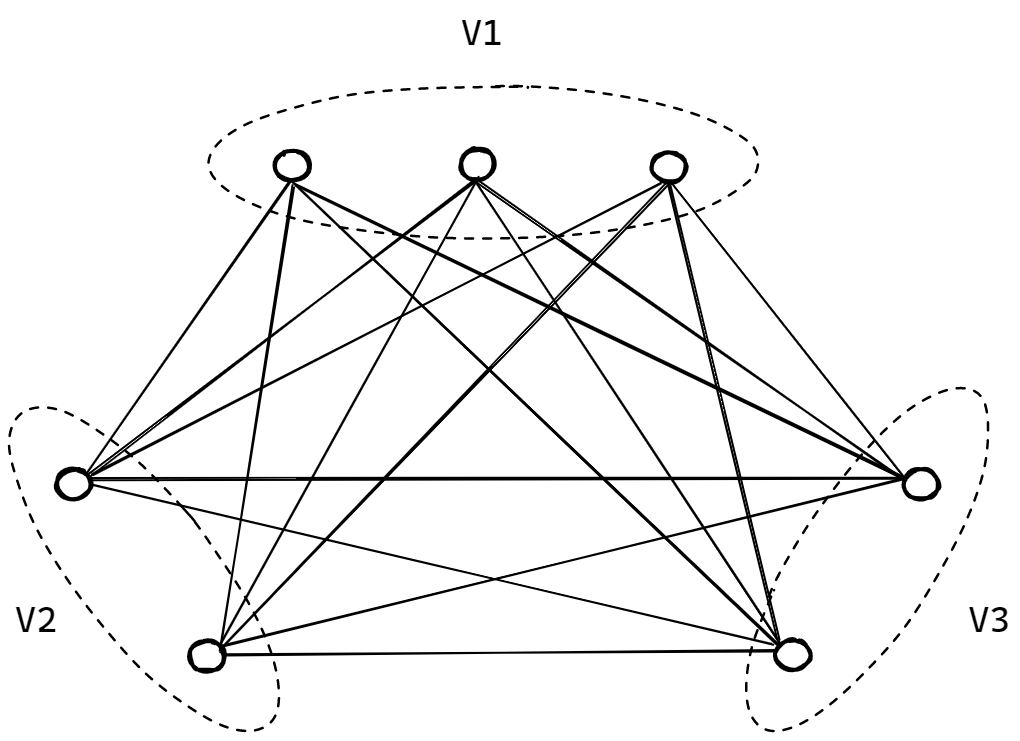
Transductive vs Inductive Learning
Embedding methods can be broadly classified as transductive and inductive. The transductive methods make an assumption that the complete graph is known beforehand. Transductive methods usually utilize both labeled and unlabeled data for training and do not generate any predictive model. Their output is an embedding lookup table. Hence these methods are also sometimes called shallow embedding methods, or encoders. To generate embedding for a newly generated node, transductive methods have to retrain the entire set of nodes. Examples of such methods include random walk based methods like DeepWalk and Node2Vec.
Inductive learning techniques are equivalent to supervised learning and do not assume the input graph to be complete or exhaustive. They are trained on training data and are applied to newly observed data outside of the training sets. As long as the newly observed data follows the same distributional assumptions as the training data, these models can generalize well to unseen data. Examples of inductive learning include Graph Neural Networks (GNNs).
Basic Graph Data Structures
Following is a list of commonly used formats for storing a graph structure, before any transformation is applied to it3.
- Adjacency matrix: a node-to-node matrix
- Incidence matrix: an edge-to-node matrix
- Edge Lists: a list of edges by their nodes
- Adjacency Lists: Lists of each node’s adjacent nodes
- Degree matrix: node-to-node matrix with degree values
- Laplacian matrix: The degree matrix minus the Adjacency Matrix (DA). Useful in spectral theory
Structural Similarity Measurement Techniques
Computing the structural similarity between nodes is a central component in graph representation learning. This similarity conveys the relative positioning of nodes in the learned embedding space. We can divide the similarity measurements for graph structures into the following two categories4.
Local Structural Similarity
Common Neighbors
The number of common neighbors for a given pair of nodes. It is calculated as the size of the intersection set of the two nodes’ neighborhoods.
$$ S(v_i, v_j) = |N(v_i) \cap N(v_j) | $$
Jaccard Coefficient
It is calculated as the normalized common neighbor value.
$$ S(v_i, v_j) = \frac{|N(v_i) \cap N(v_j)|}{|N(v_i) \cup N(v_j) |} $$
Preferential Attachment
This metric is specifically used for networks in which the number of nodes increases over time. The likelihood of a new edge associated with a node is considered proportional to the degree of the node and is denoted as:
$$ S(v_i, v_j) = |N(v_i)| |N(v_j) | $$
Adamic Index
This metric is used to calculate the similarity between two web pages based on shared features and is denoted as:
$$ S(v_i, v_j) = \sum_{v_z \in N(v_i) \cap N(v_j)} \frac{1}{\log{|N(v_z)|}} $$
Global Structural Similarity
These metrics describe the topology information of the entire network and therefore have higher computational costs.
Katz Index
This metric aggregates all the exponential paths between a given pair of nodes.
$$ S(v_i, v_j) = \sum_{i=1}^{\infty} \beta^l(A^l)_{v_i v_j}$$
where $(A^l)_{v_i v_j}$ is the adjacency matrix, $l$ is the path length, and $\beta$ is the damping factor for controlling the weights of the paths.
Rooted PageRank
This metric uses PageRank with a random walk. The similarity between two nodes can be measured as the stationary probability in a random walk.
SimRank
This metric is based on the idea that if two nodes are similar, then the nodes associated with them should also be similar.
$$ S(v_i, v_j) = c \frac{ \sum_{x=1}^{|N_{v_i}|} \sum_{y=1}^{|N_{v_j}|} S(N_x(v_i), N_y(v_j)) }{ |N(v_i)||N(v_j)| }$$
Network Embedding Models
Basic Representations
Much before the advent of deep learning, machine learning methods used to rely on graph statistics5. For example, a graph has properties like size (the number of edges), order (the number of nodes), degree distribution, connectedness, etc. A node has structural properties like degree and centrality and non-structural properties like node attributes. Centrality indicators like eigenvector centrality rank the likelihood that a node is visited on a random walk of infinite length on the graph. Betweenness centrality measures how often a node lies on the shortest path between two nodes, closeness centrality measures the average shortest path between a node and all other nodes. An edge may also have associated edge weights. Note that these features are not very helpful in designing flexible functions, and it can be very hard and time-consuming to find appropriate features for a given application. Therefore embedding methods try to leverage these graph properties as the input data.
An Encoder-Decoder Perspective
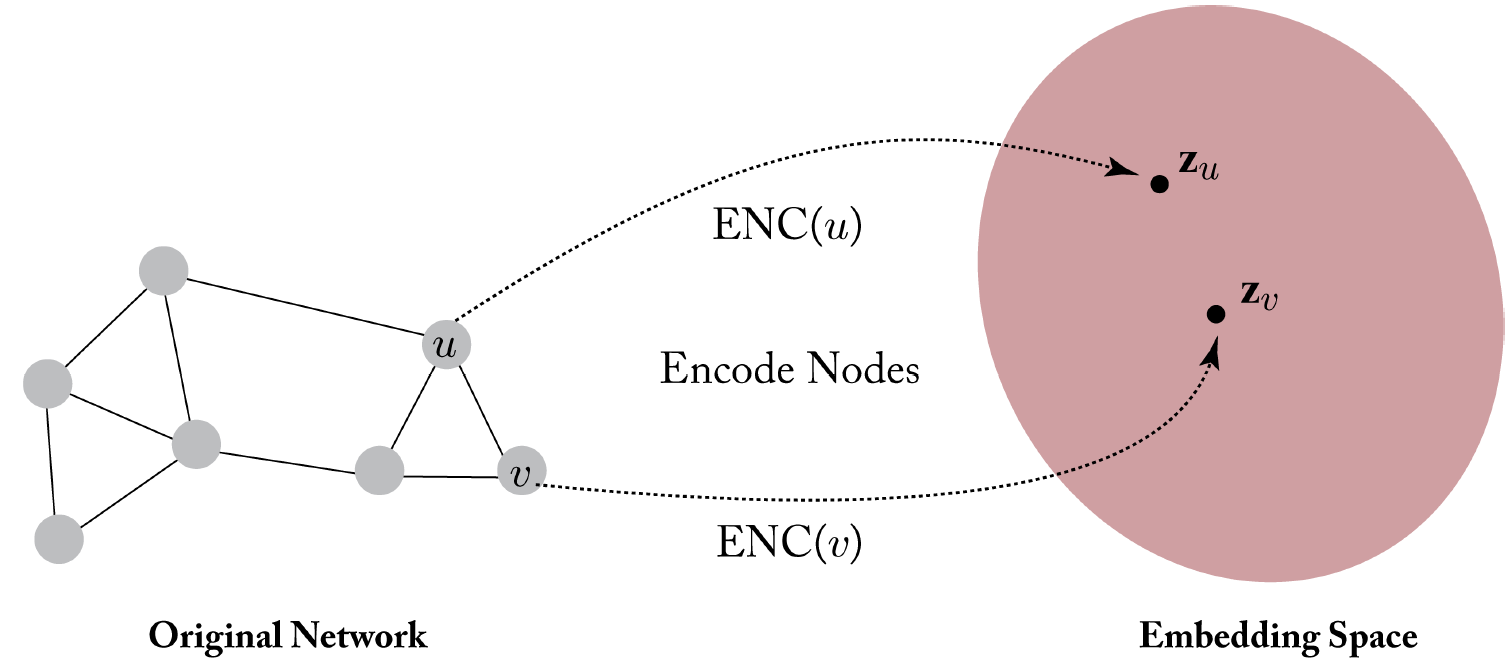
Before we dive deep into the popular modeling methods, we will build a mental model to understand these methods. Hamilton5 proposed an encoder-decoder framework for graph representation learning based on neighborhood reconstruction methods. First, an encoder maps graph nodes into a low-dimensional vector, and then a decoder uses this vector embedding to reconstruct original information about nodes’ neighbors. Overall, the system is trained using an appropriate reconstruction loss (such as mean-squared error, or cross-entropy).
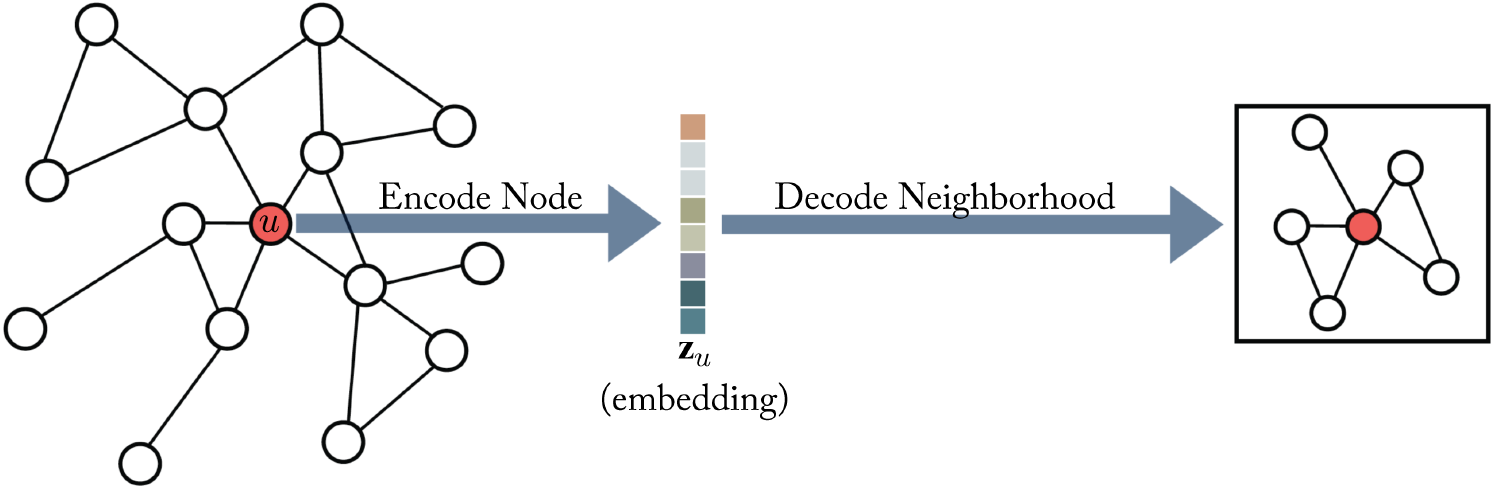
In a lot of systems, the encoder is simply a lookup table based on node IDs. Such systems are called shallow embedding methods. In Graph Neural Networks (GNNs), the encoder uses an inductive approach to generate embeddings for unseen nodes.
Factorization-based Models
As mentioned earlier, one way to represent a graph is to use its adjacency matrix, which is a square matrix whose elements indicate whether pairs of vertices are adjacent or not. This matrix can also be weighted and symmetric depending on the type of graph. Matrix-factorization based models factorize such representations to generate node embeddings.
There are many alternatives to the adjacency matrix that we can use, such as the Laplacian matrix and node transition probability matrix. First, these proximity-based matrices are constructed using the corresponding proximity measure, and then a dimensionality reduction technique is applied to generate node embeddings. Isomap, Laplacian eigenmaps, Graph Factorization, GraRep, and HOPE are among the popular methods in this category.
These methods are not very effective because the simple matrix format isn’t informative enough and can’t store complex relationships such as paths and frequent subgraphs. They are also very expensive to compute, and their high space and time complexities make them intractable for large-scale networks.
Random-walk based Models
A random walk represents a sequence of nodes where the next node in the sequence is selected based on a probabilistic distribution. There is no restriction on the walk to prevent it from backtracking or visiting the same node multiple times. The goal of this random walk is to extract rich structural information from the local structure of the graph. They try to maximize the probability that the nodes that tend to co-occur on a truncated walk should be closer in the embedding space.
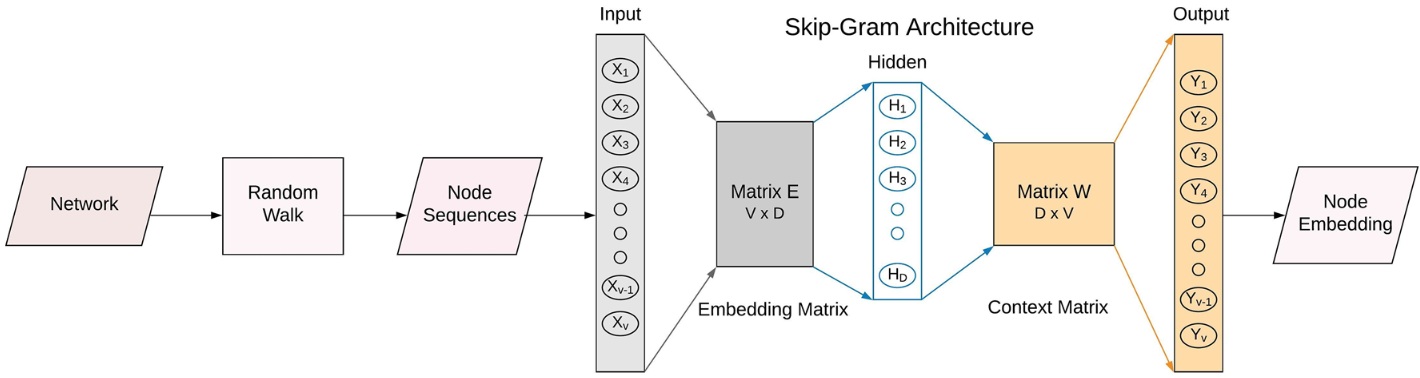
First, a truncated random walk is performed to transform graph input to node sequences. Next, an NLP-based approach is used where a skip-gram model maximizes the conditional probability of association between each input and its surrounding context. Node embeddings are learned through a shallow neural network architecture by using each node to predict its context6.
Deepwalk was one of the first models based on this idea. Node2Vec further improved it by replacing the unbiased walk (see the left figure below) with a biased walk (see the right figure below) using two parameters that control the probability of returning to the previous node and performing either a BFS or DFS.
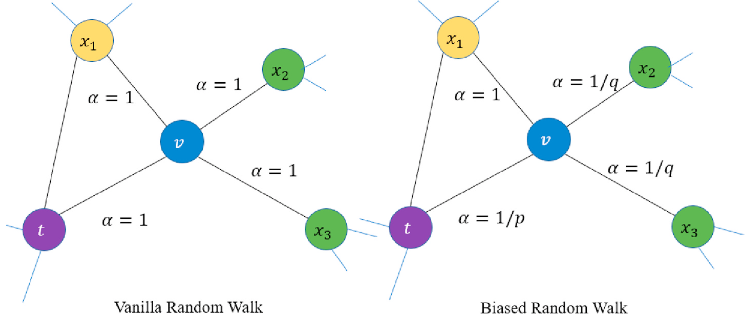
Deep Learning based Models
The methods discussed above used a piecemeal approach where a model is used to learn node embedding which is then fed to a downstream model for performing a task. Deep learning enables end-to-end solutions for these graph-based tasks and inherently learns the embedding in the architecture. Moreover, shallow neural embeddings often lead to suboptimal solutions because they can not effectively model the nonlinearity that is inherent in network data.
One naive way of using deep learning for graphs is to directly input the adjacency matrix as input to a neural network. However, this approach is not permutation invariant and depends on the arbitrary ordering of the nodes in the input matrix. Hence inductive biases must be built into the deep architectures used for graph data to effectively model this non-Euclidian, non-i.i.d. input.
There has been a surge in research on using deep learning for graph representation learning, including techniques to generalize convolutional neural networks to graph-structure data, and neural message-passing approaches inspired by belief propagation. Graph Neural Networks (GNNs) are one of the most popular deep-learning based solutions. GNNs use the Message Passing technique to encode information about the graph by successively propagating a message one hop further away in each iteration. There are diminishing returns involved when using information from multiple message-passing layers. Hence GNNs are not as deep as some of the more conventional neural networks. Different GNN models utilize different message-passing schemes. SDNE, DNGR, and VGAE are some of the popular non-GNN based deep learning models.
A Taxonomy for Graph Representation Learning
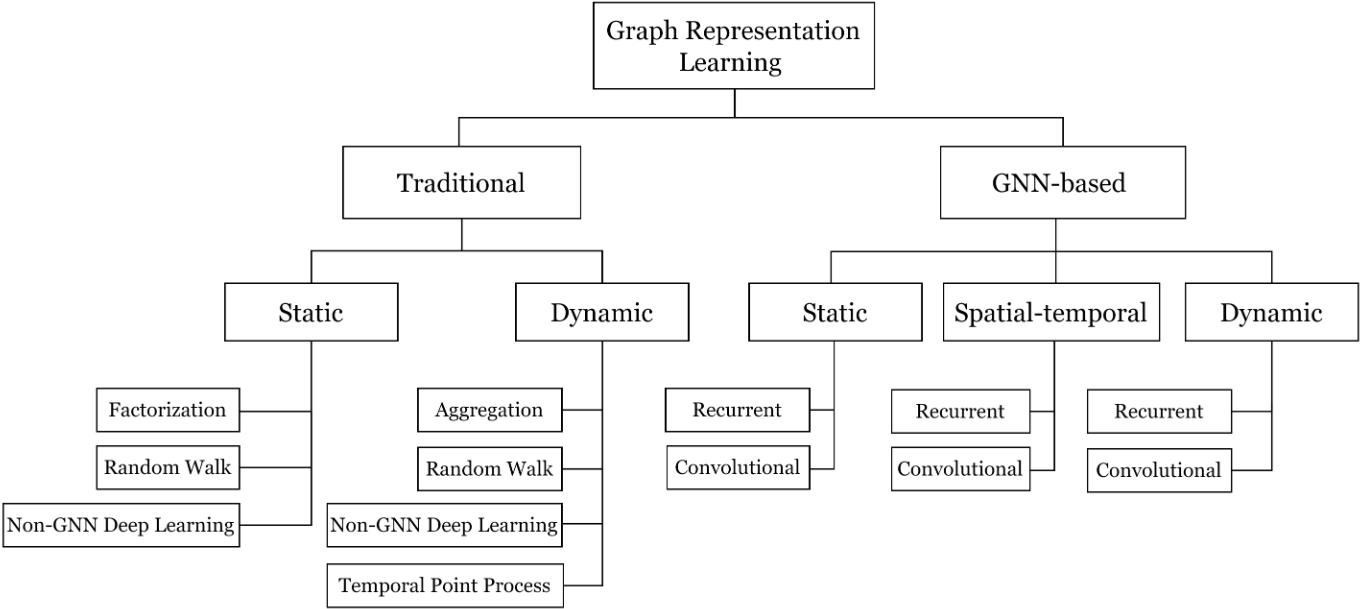
Various researchers have proposed different taxonomies to efficiently navigate the graph representation learning domain. One such intuitive approach is proposed by Khoshraftar et. al1 which first categorizes algorithms into Traditional (non-GNN) and GNN-based methods, followed by a further division into static, dynamic, and spatial-temporal graphs.
Evaluation Methods
Various downstream network analytic tasks are commonly used to evaluate learned network representations. These methods effectively measure how much of the structural information of the original graph is retained.
Network Reconstruction
We can use the learned network embeddings to reconstruct the network and examine the differences between the reconstructed and original network. precision@K and MAP metrics are often used here as evaluation metrics.
Node Classification
We can also use node embeddings as features to train a classifier on labeled nodes. This classifier can then be evaluated using the F1 score metric calculated on label predictions of randomly selected nodes from the test set.
Link Prediction
We first remove a small number of edges of the original network, and then use embeddings to predict the removed edges. Model performance on the Link Prediction task can be measured using AUC and precision@k.
Node Clustering
We can apply k-means clustering on the learned embeddings and evaluate the detected communities. Metrics like accuracy and NMI can be used in this situation.
Visualization
The learned embedding space can be reduced to 2 or 3 dimensions by applying t-SNE or PCA. For good-quality embeddings, we should be able to see the nodes belonging to the same class or community clustered together.
Applications of Graph Embedding
The insights learned from the complex graph structures can be useful for various scientific and business use cases. This section highlights some of the applications4.
-
Community Detection: Representation learned from network structure and node attributes enables the identification of meaningful communities in social networks.
-
Recommendations/Targeted Advertising: Graph learning based recommender systems are used, for example, in social networks to recommend friends.
-
Privacy Protection: The degree of trust between two nodes can be estimated using their embeddings which can be used to control the sharing of private information in social networks.
-
Network Visualization: Low-dimensional vector representations make it possible to use methods like PCA and t-SNE to effectively visualize large graphs.
-
Network Alignment: Graph representation learning can help in connecting multiple graphs in the same system by identifying common nodes across different networks.
-
Network Compression/Summarization: Vector embeddings can also be used to describe the input network with a compact representation while also having the capability to effectively reconstruct the network.
Advanced Topics
The vast majority of the literature in graph representation learning is focused on non-attributed, static, and unsigned homogeneous networks. This means that there are several challenges that need more extensive research when applying these techniques to real-world graphs. For example, the propagation of information between nodes and links in heterogeneous networks is much more complex than the propagation in homogeneous networks. Applying graph learning to enterprise graphs with billions of nodes and edges is also very complex. Fortunately, various proposals have been made to learn effective graph representations from heterogeneous and dynamic networks. And, many recent research works have proposed adjustments to the training and inference algorithms, and storage techniques to enable learning from large-scale graphs. In the coming set of articles, I will highlight some of the most important research on this theme.
Summary
This article presented a comprehensive review of graph representation learning. In the upcoming series of articles, I will dive deep into individual modeling techniques. I will also highlight the current state-of-the-art methods and their real-world applications in different industries. Additionally, I will focus on optimizations that enable large-scale usage of these methods, along with articles on graph representation learning usage for recommender systems.
References
-
Khoshraftar, S., & An, A. (2022). A Survey on Graph Representation Learning Methods. ArXiv. /abs/2204.01855 ↩︎ ↩︎
-
Zhang, D., Yin, J., Zhu, X., & Zhang, C. (2017). Network Representation Learning: A Survey. ArXiv. /abs/1801.05852 ↩︎
-
Broadwater, K. (2023). Graph Neural Networks in action. S.l.: O’Reilly Media. ↩︎
-
Luo, Yu, Sai, Cai, Cheng. (2022). A Survey of Structural Representation Learning for Social Networks. Neurocomputing. 496. 10.1016/j.neucom.2022.04.128. ↩︎ ↩︎
-
Hamilton, W. L. (2020). Graph representation learning. San Rafael, CA: Morgan & Claypool. ↩︎ ↩︎
-
Mohan, A., & Pramod, K.V. (2019). Network representation learning: models, methods and applications. SN Applied Sciences, 1. ↩︎
Related Content
Did you find this article helpful?